With the increasing access to exome and genome testing the interpretation of this data is taking longer and longer, as even the most highly skilled and experienced individuals, cannot be an expert on every gene they encounter.
As labs need more time to familiarize with previously un-encountered genes and variants, the time taken to complete cases has been increasing. With a limited global workforce of skilled professionals and limited time for analysis this is leaving many with a lack of resource
to dedicate to complex cases, leading to a risk of reduced diagnoses for patients.
Recurring “known” pathogenic and likely pathogenic variants are reported in a wide range of disorders. Thanks to smart automation capabilities, using the Congenica clinical decision support software you can now automatically pre-classify known variants observed in your rare disease cases, significantly saving time to maximize case throughput and improve your diagnostic yield and outcomes for patients.

When we talk about automation, what do we mean?
The goal of automation is to accurately and safely process a sample from data to report, identifying any pathogenic variants without requiring human input for repetitive tasks.
Automation can reduce the time to resolve a case in two ways: reducing the number of variants for review, by automatically processing all previously known variants; and reducing the time spent reviewing each variant, by providing high confidence in-silico predictors or automatic application of ACMG criteria and classification.
The basics of the sample processing workflow are the same the world over. From data upload, familiarizing yourself with the background of the case; to analyses, filtering your data to remove unlikely candidates, before interpretation and reporting.
Congenica is automating this process in two ways:
- automating known variants of clinical interest and
- automatically adding additional evidence for all variants of interest.
Our dual strategies address the challenges around limited clinical genetic resources and incomplete knowledge of genomic variants. We call this, Congenica Express.
Using Congenica Express you can now automate the interpretation of variants described in your previous case-sets or in curated databases. The Congenica platform can prepopulate variant classifications, ACMG criteria, supporting comments, relevant literature articles and report-ready summary statements, using pre-curated information provided by your selected datasets. This means less duplicated work and more rapid analysis for recurrent variants.
Congenica Express combines user curated data with case-relevant gene panels to automatically interpret variants when they are seen again in a future individual.
-- Helen Savage, DipRCPath, Lead Clinical Scientist, Product Innovation
Congenica comes with key benefits for automation
We make it easy for you to get started. You can accelerate your turnaround times, increase case throughput, improve safety and quality, and maximize diagnostic yield.
- Automate your workflows
- Benefit from pre-curated variants lists, gene panels for each case and a custom report template.
- Achieve 5-minute interpretation times
- In internal testing, Congenica’s automated classification of known variants significantly reduces time to report – achieving complex genomic data interpretation in as little as 5 minutes.
- Increase your case throughput
- Congenica software already enables the fastest analysis and highest
throughput of complex genomic cases. With Congenica Express, labs can increase their throughput by a further 20% to 60%, depending on the frequency of recurrent variants. [1]
- Congenica software already enables the fastest analysis and highest
- Maximize your diagnostic yield
- Compared to industry averages, the Congenica decision support software enables laboratories to reach actionable insights from their analysis in 30% more cases. [2]
- Congenica Express will detect and classify previously curated variants present in your chosen dataset. It will pick out the “easy wins” and highlight them for you to avoid repetitive work.
- Have complete confidence in the quality of your automated analysis
- Congenica Express features built-in functionality to safeguard against issues such as false negative reporting and unmasking carrier status in unconsented individuals.
Congenica enables us to analyze variants easily and quickly,
saving a lot of time in our NGS data.
-- Dr Youngmok Lee, Assistant Professor, UConn Health, USA
Congenica Express improves the flow through a decision support system
The current flow through any decision support system such as Congenica includes the
following standard steps:
- Review of the background to the case
- Review the quality of the data provided for analysis
- Variant analysis and interpretation
- Second check the results, to avoid false negative or false positive reporting, before completing case
- Authorize and report the results
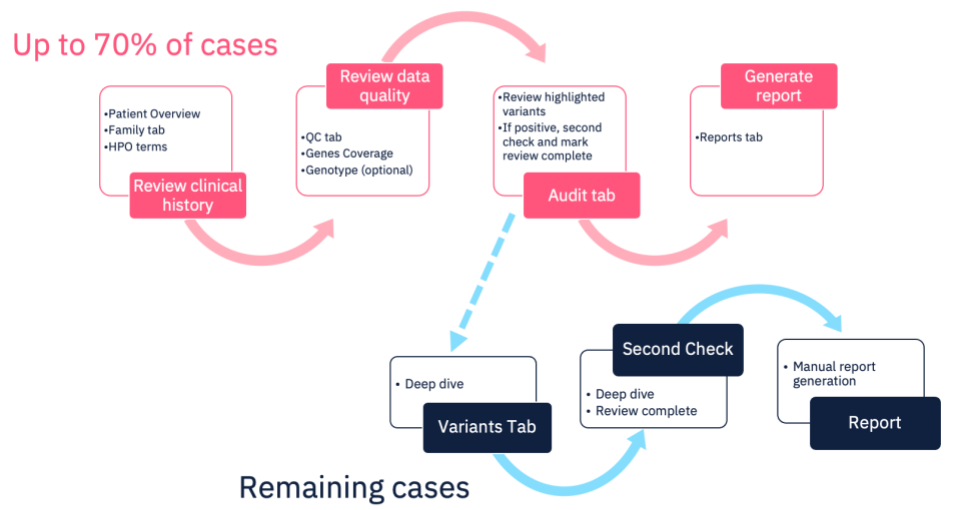
Currently in the Congenica system this is the process followed for all cases. However, using our Express pipeline, up to 70% of cases will not need a “deep-dive” as our pipeline will interpret and report any curated variants of interest without the need for human intervention – presenting the interpretation findings for review and confirmation. Of course, if no causal variants are identified in the pre-curated data, that doesn’t have to be the end – the case can be analyzed using our existing functionality, allowing you review and interpret any novel variants, according to your SOPs.
Automated Analysis with Congenica Express
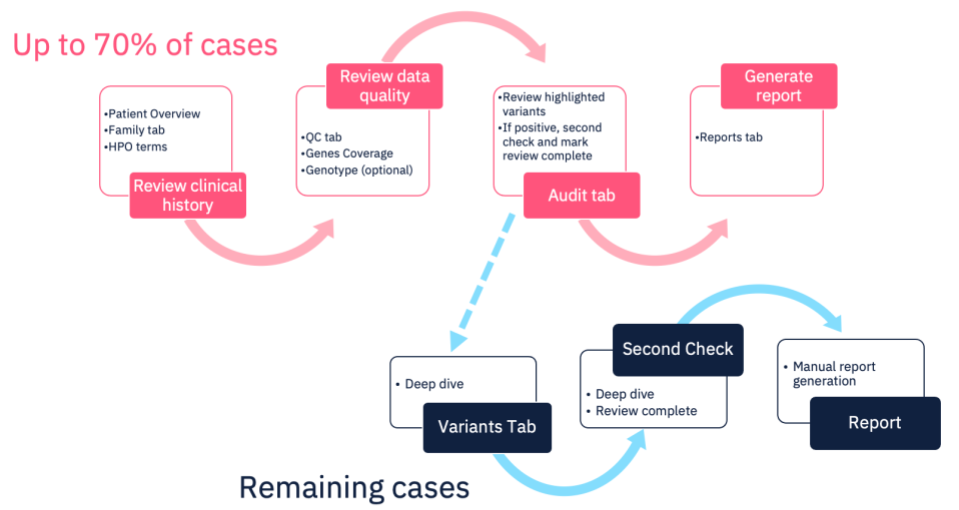
Use cases
Automating the interpretation of previously classified variants has huge potential across multiple applications and use cases, for example, to provide a “pre-screen” of rare disease cases to quickly identify, interpret and report on variants previously classified, with
confidence.
Recurring “known” pathogenic and likely pathogenic variants are reported in a wide range of disorders, including:
- >70% of pathogenic variants in cardiomyopathy patients [4]
- 35% of pathogenic variants in cardiac arrhythmia patients [4]
- 50% of de novo missense variants in neurodevelopmental patients [5]
- Recurrent PAX6 variants resulting in ophthalmic malformations in 15 families[6]
- 15,600 Pathogenic or Likely Pathogenic SNVs and indels in ClinVar reported in multiple individuals with no conflicting interpretations
If this pre-screen does not detect any previously classified variants, customers can continue to perform a deep dive into the novel variants detected in an individual.
Case Study: Driving down turnaround times and costs
In a study analyzing 19,000 cases from the Congenica knowledgebase, a team of registered clinical scientists identified nearly 4,000 cases with recurrent pathogenic or likely pathogenic variants, which Congenica Express of Known Variants would have automatically classified, increasing throughput significantly.
Using the Congenica Automated Classification of Known Variants pipeline, interpretation and reporting of each of these 4,000 cases could be completed in under 8 minutes, saving:
- almost 5,500 hours (91%) of combined interpretation effort
- and 98% of staff costs!
Find out more
Congenica Express achieves complex genomic data interpretation in as little as 5 minutes when a recurring causal variant is identified, maximizing lab throughput.
Learn more about Congenica Express
References
[1] White Paper: Analyze, Interpret and Report NGS Data Faster than Ever Before –
https://www.congenica.com/efficiency
[2] Genet Med. (2019) 21: 3–16. Also see J. Med. Genet. (2019) 0, 1 – 9. Automating Clinical Genomic Analysis and Reporting for Rapid NGS
[3] Webinar: Accelerating the Identification of Genetic Diseases –
https://www.congenica.com/accelerating-the-identification-of-genetic-diseases/
[4] Dutch founder variants in cardiac disease –
https://link.springer.com/article/10.1007/s12471-019-1250-5
[5] Neurodevelopmental disorders –
https://europepmc.org/article/MED/29100083
[6] Ophthalmology –
https://pubmed.ncbi.nlm.nih.gov/31700164/
[7] Congenica Express™ data sheet
https://genomics.congenica.com/hubfs/02%20Datasheets/Congenica%20Automation%20Datasheet.pdf



.png?width=320&height=192&name=Add%20a%20title%20(2).png)
