With case loads and the complexity of genomic medicine increasing every day, clinical teams are experiencing increasing pressure to make their variant interpretation more efficient. However, for many laboratories, challenges exist that reduce efficiency, such as the processes for data submissions, or limitations in the prioritization of variants, slowing interpretation.
At Congenica, we always work to enable our partners to find life-changing answers for patients more quickly and confidently. We are therefore excited to highlight a number of new enhancements to our software in Congenica v2.4 that will overcome these challenges.
Updates include a new in-silico deleteriousness score, CADD, to inform clinical interpretations and an overhaul of our data submission tool to enable more flexible and streamlined ways of working for sequencing and clinical interpretation teams.
Data submission tools designed for your workflows
Often, clinical geneticists will work separately from the technicians or service providers who sequence samples from patients. This can result in logistical and communication challenges when trying to submit variant interpretation requests (IRs) and sequence data files simultaneously. Labs will often need to setup their own secure file transfer systems and protocols, costing them time and money, with slow connections & interrupted uploads delaying the time from sample to diagnostic report. Clinical teams needed to coordinate with and rely on limited bioinformatics resources to upload IRs and samples creating bottlenecks that can delay diagnostic answers.
Congenica’s new data submission tool allows customers to upload genetic data files and create/upload Interpretation Requests separately and at different times (up to a week apart). To make this easier, we have also upgraded our Interpretation Submission Request page to enable non-technical users to upload or create an IR through a simple graphical interface.
We’ve removed the headache of supporting your own servers or relying solely on command line tools to upload files by hosting our own secure FTP (sFTP) server for uploading FASTQ, BAM and VCF_SNV files. This can be accessed securely by lab technicians, regardless of whether they are users of your Congenica platform.
What all this means is that you can easily and securely adapt Congenica to your lab’s existing SOPs, whether you have separate sequencing and interpretation teams in-house or out-source sequencing to third-parties, while enabling non-technical clinical teams to submit interpretation requests.
New data supporting your interpretation confidence
Often, in-silico pathogenic scores play a key role in “sense-checking” a variant interpretation. However, many in-silico scoring tools only search for possible missense mutations caused by SNVs and indel variants.
Congenica has now integrated in-silco CADD scores (Combined Annotation Dependent Depletion) into the platform. Unlike other in-silico tools, CADD scoring integrates dozens of genomic features to predict the information about variants (PMID: 3037182). By making this available to users worldwide, we’re providing another source of confidence for clinical diagnosis.
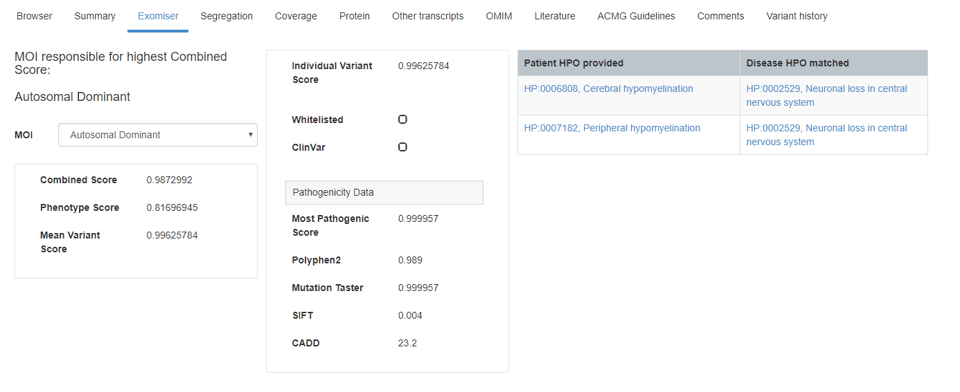
CADD scores are displayed in the new Exomiser tab, which also provides transparency into how Congenica uses mode of inheritance, HPO symptom-to-variant matching, and in-silico scoring to prioritize variants for your interpretation. The Exomiser tab also puts all of your in-silico scores in one place, for a quick understanding of a variant’s possible pathogenicity.
We appreciate your feedback and the support from our partners in the genomic medicine community more than ever during these challenging times as we stay committed to our goal of enabling genomic medicine.
If you have any questions about using the improvements mentioned in this blog post or would like a copy of the complete release notes, contact support@congenica.com.





