Didn’t attend ESHG 2021? Don’t worry, this blog enables you to catch up on all the Congenica content that you missed.
Satellite session: Accelerating NGS analysis: automating best clinical practice to provide better outcomes for rare disease patients
The increasing demand for complex genomic data interpretation and reporting services creates a growing challenge for laboratories. Lengthier analysis required for complex rare disease analysis is limiting case throughput and delaying results for patients.
In this session, leading European institutions share how clinical decision support software is enabling them to provide life-changing answers for more patients, faster than ever before, by replicating best clinical practice and automating many of the manual processes in the interpretation and reporting of NGS data.
- The Institute of Immunology and Genetics (IIG) discusses postnatal and prenatal whole exome cases, highlighting how they are able to achieve incredibly short turnaround times to benefit their patients.
- The Tettamanti Research Center presents pediatric case studies to demonstrate how they have significantly reduced the time it takes, from initial patient referral by a clinician to delivery of results.
The session also details some of the latest developments in next generation genomic data analysis, enabling data interpretation with the highest efficiency, accuracy and confidence.
Poster: Exome analysis of prenatal and postnatal cases referred with skeletal dysplasia: an overview of genomic and phenotypic findings.
Awarded ‘best poster’ at ESHG 2021
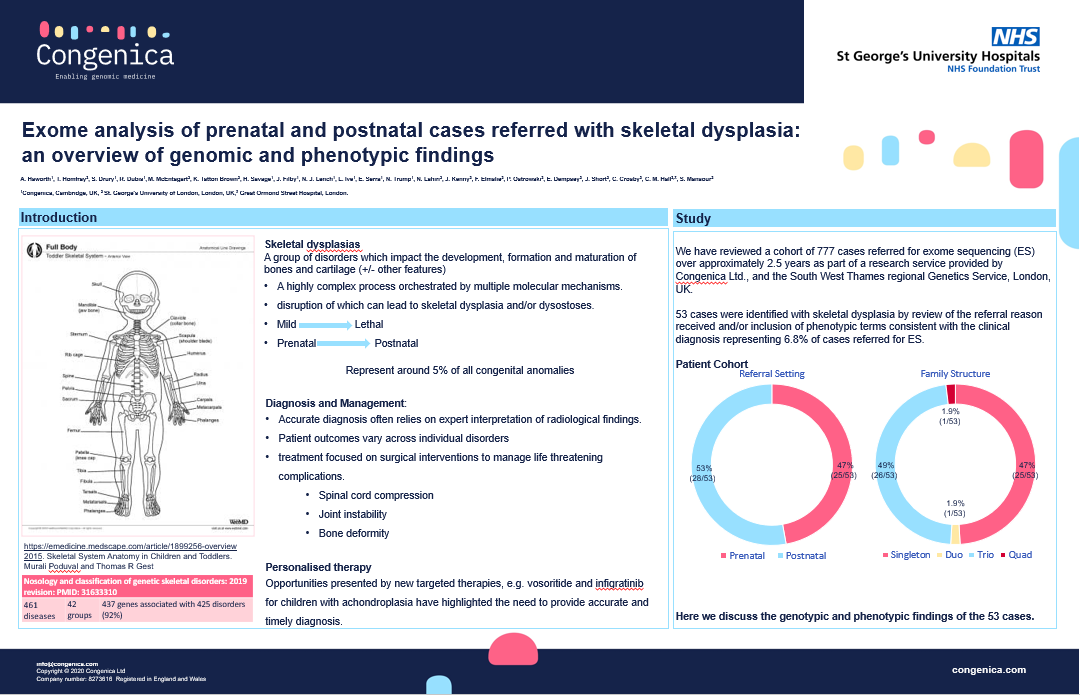

Skeletal dysplasias (SD) are rare disorders representing approximately 5% of all congenital anomalies. They are highly heterogeneous and clinical findings are often non-specific, so accurate diagnosis often relies on expert interpretation of radiological findings.
In this poster, Andrea Haworth, lead clinical scientist, rare disease, Congenica, describes the genomic and phenotypic findings of SD cases referred to clinical genetics over an approximately 2.5-year period.
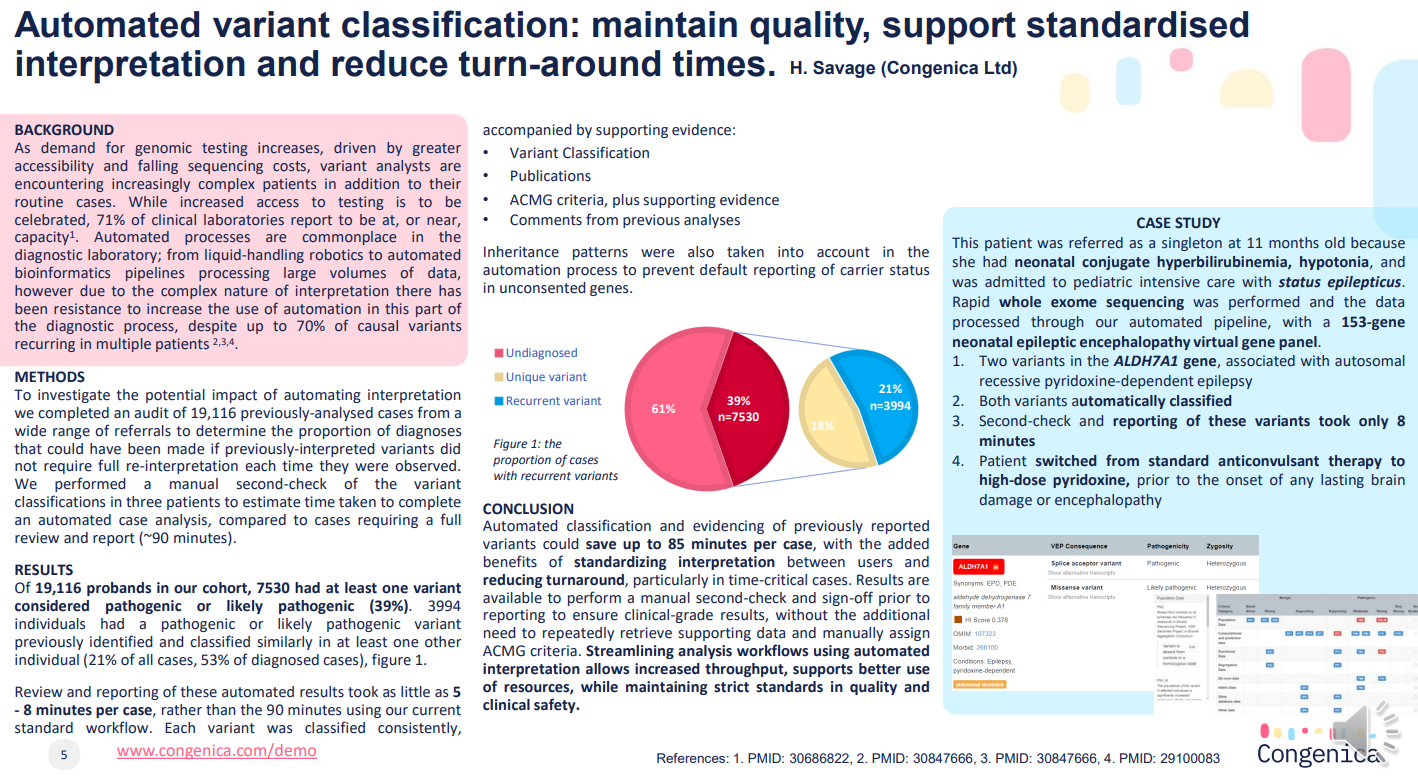
Poster: Automated variant classification workflows maintain quality standards, support standardisation and reduce turn-around times in a rare disease laboratory
As demand for genomic testing increases, variant analysts encounter increasingly complex patients in addition to their routine cases. While increased access to testing should be celebrated, the use of first-line exome and genome testing makes variant interpretation a key bottleneck, as highly skilled analysts are not experts on all genes/disorders encountered
In this poster, Helen Savage, lead clinical scientist, product innovation, Congenica, presents the case for automating analysis of cases, to rapidly generate high-quality, relevant results supporting the diagnosis of patients with rare disease, without compromising the diagnostic yield each analysis.

.png?width=674&name=Enabling%20Genomic%20Medicine%20Best%20practice%20in%20genomic%20analysis%20(2).png)



.png?width=320&height=192&name=Untitled%20design%20(8).png)
.png?width=320&height=192&name=Since%202016%2c%20the%20number%20of%20women%20working%20in%20STEM%20fields%20in%20the%20UK%20has%20increased%20by%20216%2c552%2c%20taking%20the%20total%20number%20over%20the%201%20million%20mark%20for%20the%20first%20time.%20Women%20now%20make%20up%2024%25%20of%20the%20STEM%20workforce%20i%20(2).png)
-1.png?width=320&height=192&name=Deciphering%20Developmental%20Disorders%20(1)-1.png)